Through several recent interactions I’ve been reminded just how much of the information I take for granted about the state of genomic resources for different grass species, the relationships between different grass species and even the correspondence between common and scientific names is the product of my own meandering career path to date and doesn’t, by any stretch of the imagination, represent common knowledge, even among plant biologists.
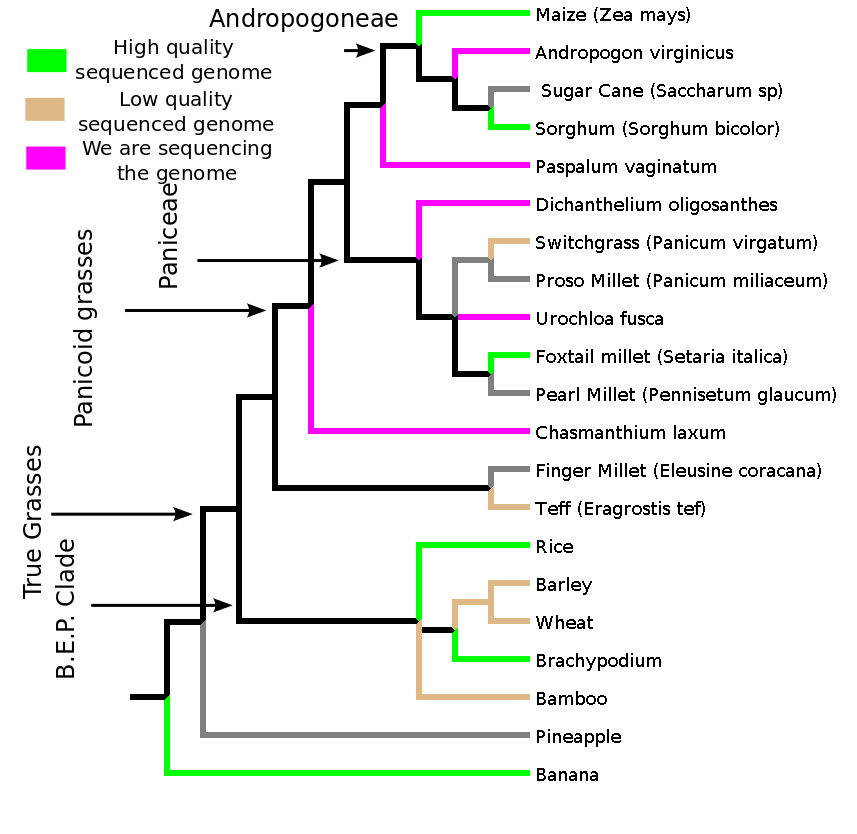
High and low quality genome assemblies are here defined as assembles with pseudomolecules (essentially chromosomes) that contain most of the core grass gene complement, and assemblies either without pseudomolecules, or where a large fraction of genes aren’t placed on the pseudomolecules yet. “We” in the legend above refers to a project from JGI’s Community Science Program (CSP).
This is by no means a comprehensive treatment. There an estimated 11,000 grass species, the vast majority of which are known only from physical observation (not so much as a single fragment of their dna has ever been sequenced).
However, if you’re interested in a more comprehensive treatment than my off the top of my head approach, go here and download “NPH_3972_sm_FigS1-S2.pdf” to see a tree of more than 500 total grass species, representing pretty much every single major group within the grasses. It’s my go to reference whenever I’m trying to discover the relationships between grass species I’m not familiar with and as far as I know it’s still THE reference work to which all other grass phylogenetics papers are compared (but feel free to let me know in the comments if you know of a newer paper that includes even more species).
[…] Amaze your friends with you up-to-date knoweldge of the current state of grass genomics. […]
Pingback by Nibbles: Old basil, Old newsletters, New old vegetables, New network, New phylogenies, Old story — June 10, 2015 @ 12:11 am