Thinking about defining the number of genes present in the maize genome reminded me of an old* story about the trouble of defining what truly represents a gene and how really awesome ideas can sometimes come years before the data needed to support them.
The year is 2002. The first complete version of the human genome is still a year away. The genomes of two plant species have already been published (rice and arabidopsis) but in terms of shere genome size, both species are a drop in the bucket compared to the human genome, or other plant genomes like corn or wheat. But none of this is particularly important except to set the stage.
Two researchers at Rutgers University were sequencing a tiny piece of the maize genome (~0.01%) that surrounded a single gene call bronze1 — the fifth most studied gene in maize — when they found something unexpected.
They had previously 10 identified genes in a single stretch of 32-kb of the maize genome. (A similar gene density throughout the remainder of the maize genome would have resulted in a maize genome containing more than 700,000 genes!) However it was already known that the maize genome was split between small gene-rich islands and vast desolate expanses of transposons (referred to as transposon nests**), and in fact the same study identified a couple of these nests of transposons on either side of their gene rich island (see part A of the second picture in this post).
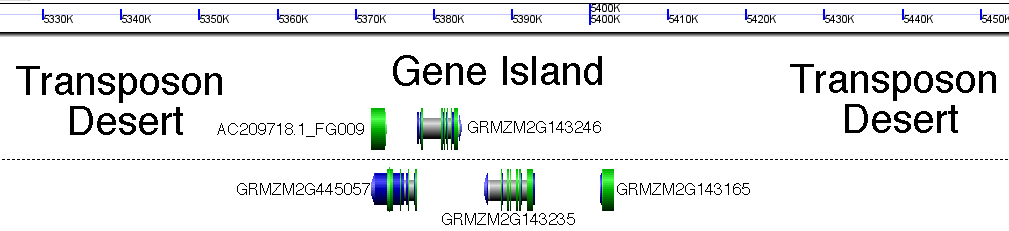
Below I'll use cartoons, but here's a real and to scale example of a gene rich island I picked at random from maize chromosome 3. Genes and intergenic spaces are to scale. Base image generated with GenomeViewer, part of the CoGe toolkit. http://www.genomevolution.org/CoGe/
Their initial sequencing used DNA from a breed of corn called McC, which I must admit I’ve only ever read about in this particular paper. However, when they decided to sequenced the same region from the genome of B73*** they made three discoveries which I’ve listed in increasing order of strangeness:
- The giant nests of transposons on either side of the bronze1 gene island were made up of completely different transposons in the two varieties of corn. While this was surprising, transposons are generally squirrelly pieces of DNA. (Part B of the image below.)
- An new group of transposons had split the bronze1 gene island apart in B73 (Part C of the image below).
- The last four genes of the island were completely missing (Part D of the image below).
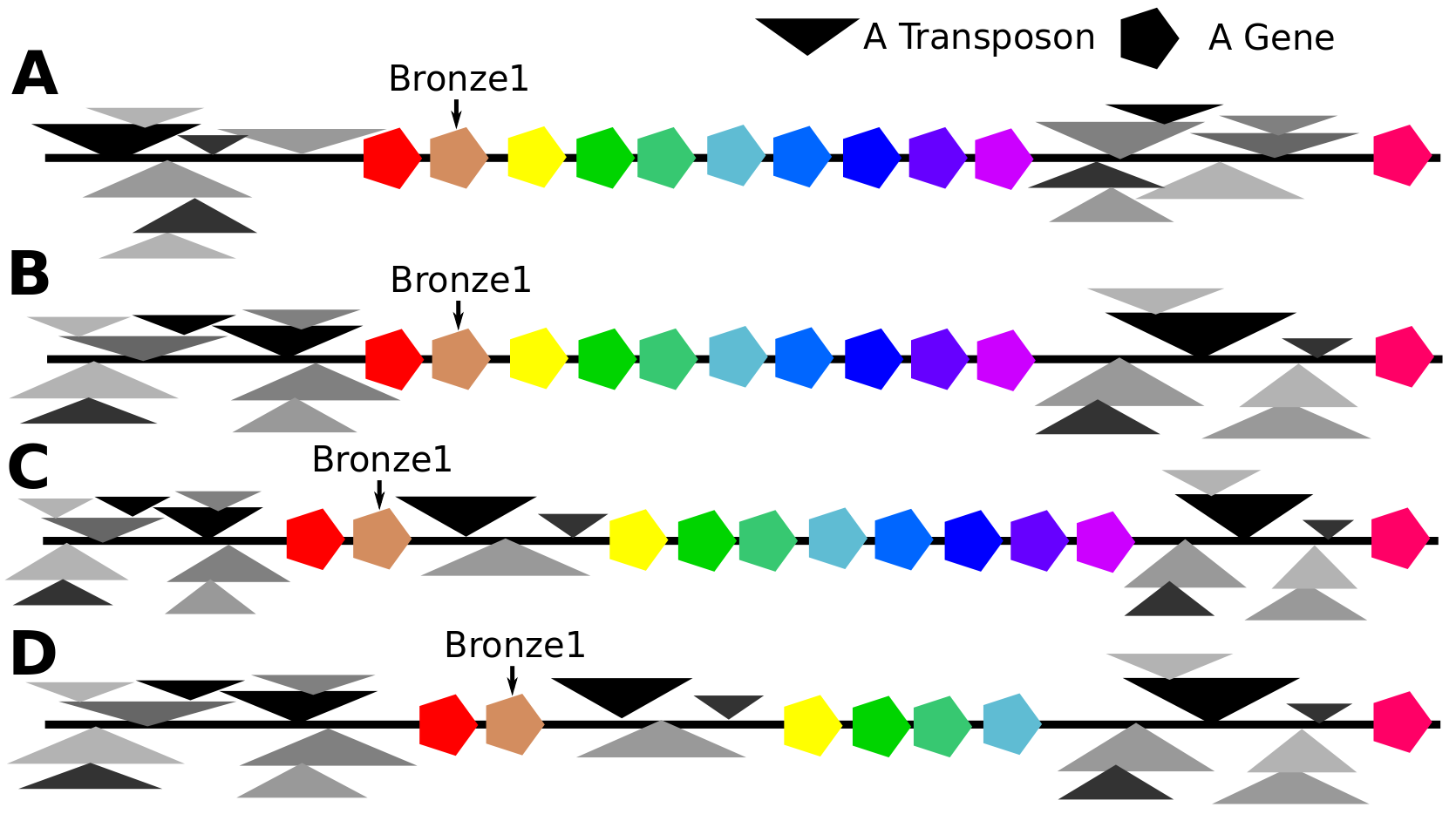
A cartoon summarizing the differences between the bronze1 gene island in the inbred McC (part A) and the same region in B73 (part D), the kind of corn that was used in the maize genome sequencing project. Part B shows the completely different set of transposons found in the two transposon nests bordering the bronze1 island. Part C shows the new cluster of transposons found between genes in the bronze1 island, and part D shows the disappearance of four genes from the bronze1 island. Absolutely nothing is to scale. The arrangement of transposons within clusters in this image was completely random.
The last was a finding that had enormous implications. Out of 13 genes (the ten in the island, plus three more on the far side of one of the transposon nests), four were completely absent in a comparison between two inbred lines. It was a finding that had huge implications for explaining heterosis or hybrid vigor — the observation that the offspring of dissimilar parents tend to be stronger and healthier than the offspring of closely related parents.
The Big Idea:
If you’re a corn plant that is missing a bunch of genes, it’s a good bet your relatives are missing a lot of the same genes, so if you reproduce with them, your kids will have the same missing genes (and the same problems) that you do. But when you mate with a really distantly related corn plant, even if its also missing a lot of genes, they’re not likely to be the same genes, so your children will have at least one working copy of most of the genes either of their parents were missing, and be stronger and healthier as a result.
But there’s a two part punchline to this story:
-The first part is that the lost genes reported in this paper weren’t really genes at all but fragments of genes captured by another type of transposon called helitrons. (A finding reported in a paper from the same lab that made the original discovery (2).) As helitrons move around the genome, they can copy pieces of real genes and carry them with them to their new location. Because the gene fragments are copies of actual genes, they can be dead ringers for the real thing, and the helitron and their capture gene fragments contribute significantly the the difficulty of estimating the true number of genes present in the genome of corn, even a year after the publication of the corn genome. Helitrons also have important implications for evolution, since they provide a mechanism to mix and match pieces of existing genes, potentially creating an occasional new and evolutionary beneficial gene.
-The second part of the punchline is even better. Even though the original lost genes of the bronze1 gene island turned out to not be real genes at all, the arguments made in the original paper for how the complete absence of genes from some inbreds could contribute to hybrid vigor have taken on a new life recently in the maize (corn) genetics community, after the publication of a paper this October that found more than 10% of the genes we’re most confident truly are genes are missing from the genomes of some kinds of corn (3). So even though the original data used to support the idea turned out to not support it after all, the idea that restoring copies of lost genes is at least part of the explanation for hybrid vigor may still prevail. (It has be conclusively shown that the gene content of different kinds of corn really are different, but someone still needs prove that restoring copies of these missing genes makes hybrid plants bigger and stronger).
1. Fu H, & Dooner HK (2002). Intraspecific violation of genetic colinearity and its implications in maize. Proceedings of the National Academy of Sciences of the United States of America, 99 (14), 9573-8 PMID: 12060715
2. Lai J, Li Y, Messing J, & Dooner HK (2005). Gene movement by Helitron transposons contributes to the haplotype variability of maize. Proceedings of the National Academy of Sciences of the United States of America, 102 (25), 9068-73 PMID: 15951422
3. Swanson-Wagner RA, Eichten SR, Kumari S, Tiffin P, Stein JC, Ware D, & Springer NM (2010). Pervasive gene content variation and copy number variation in maize and its undomesticated progenitor. Genome research PMID: 21036921
*Old by my standards, since I hadn’t even begun to study biology back them.
**When some varieties of transposons jump from one location to another in the genome, they prefer to land inside DNA that codes for other transposons instead of near genes. So one transposon lands in between two genes by chance, more transposons target it as a safe place to insert, and before you know it (relatively speaking) a cluster of transposons 100 kilobases or longer has wedged itself into the genome.
***The variety of corn that would ultimately be the source of the complete maize genome seven years later.
[…] This post was mentioned on Twitter by Shrikant Mantri, James. James said: The idea that hybrid vigor might result from missing genes is an old one, with a couple plot twists. A J+TGC post: http://bit.ly/hppE8n […]
Pingback by Tweets that mention Hybrid vigor and missing genes – James and the Giant Corn -- Topsy.com — November 27, 2010 @ 8:53 pm