Colored aleurone1 and Purple plant1 are both genes with long histories in maize research and are involved in the regulation of anthocyanin biosynthesis.The mutant version of purple plant1 does exactly what it sounds like. (In the proper genetic background) it has plants producing anthocyanin (a purple plant pigment) everywhere, resulting in purple plants. The mutant form of colored aleurone1 was identified from a mutant that changed the color of individual corn kernels. Guess which of these two classic maize mutants made it into the top 15 most published on genes in maize, and which fell barely short.
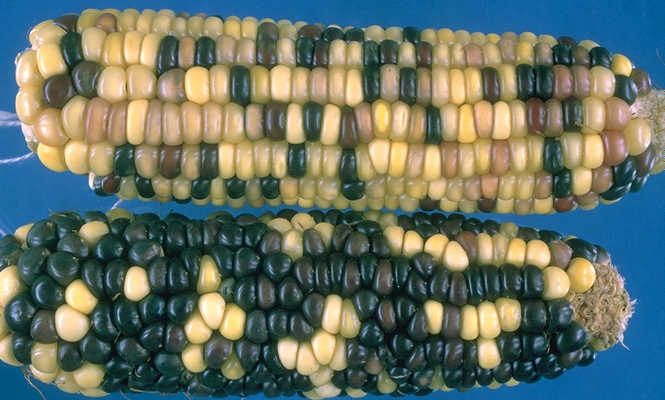

Ears segregating for the colored aleurone mutant phenotype. Image courtesy of MG Neuffer via MaizeGDB.
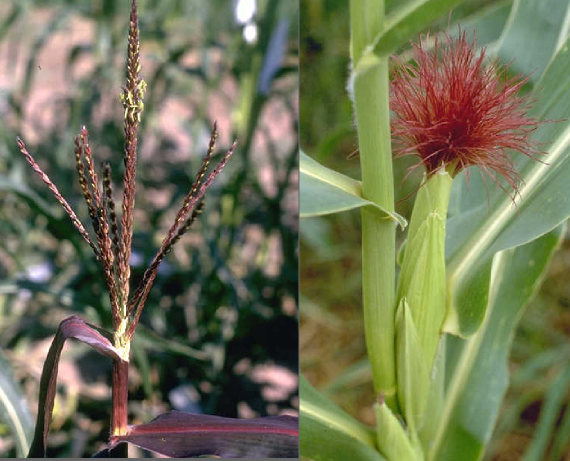

Purple plant1's phenotype is highly variable depending on the genetic background the mutant is in. Images courtesy of MG Neuffer via MaizeGDB.
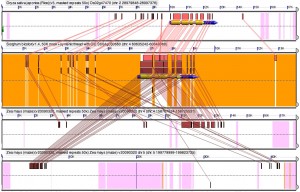
The two genes are also duplicates (homeologs) resulting from the maize whole genome duplication. From the picture below you can also see both the two genes and the regions they are in match up to single regions in rice and sorghum, two grasses that haven’t gone though a whole genome duplication since the great radiation of grass species that took place an estimated 50 million years ago (well after dinosaurs stopped walking the earth). (more…)